

Document Type : Original Article
Authors
1 Department of Biotic Evolution- Marine Science Center, Basrah University, Basrah, Iraq
2 Department of Biology, Collage of Education for Pure Science, Basrah University, Basrah, Iraq
Abstract
Keywords
Chemotaxis is the mechanism by which a single-celled organisms, bacteria and somatic cells direct their activity toward compounds in their own surrounding environment. Chemotaxis is considered positive when a movement towards the used compounds occurs, such as in the movement of bacteria towards amino acids, sugars, bilateral peptides, and purines, while it is considered negative when movement is in the opposite direction, as in bacteria away from antibiotics, toxins and some alcohols and fatty acids [1].
[2] stated that chemotaxis plays an important role in the development of biofilms as the bacteria produce chemical compounds that serve as an attraction agents between the bacteria and adhesion surface, while in the later stages of biofilm, other chemicals contribute to the destruction of the biofilm and thus moving away from the surface of adhesion [3]. Previously, [3][4][5] reported that bacteria with positive chemotaxis can travel to suitable sites for survival and growth in complex environments and that nutrient molecules (amino acids and sugars) are not all considered attractive, while on the other hand, not all attractants are good nutrients.
Microbial secondary metabolites include a wide range of naturally produced compounds that perform a large number of vital activities. Secondary metabolites are not involved in growth, development, or reproduction; but support microbes living in its environment [6]. Secondary metabolites also play an important role in the physiological adaptation and many of them inhibit the growth of competing microbes [7].
Bacterial species belonging to the genus Bacillus can produce many secondary metabolites with antimicrobial effects towards other bacteria such as lipopeptides, polypeptides, macrolactones, fatty acids, polyketides, and isocumarines [8]. Therefore, these bacterial species were widely used in the medical and pharmaceutical industries for commercial production of antimicrobial substances [9][10] .
Several studies have been carried out to extract secondary metabolites of Bacillus species to investigate the inhibitory effect of these extracts against many plant and animal pathogens and also against biofilm-forming bacteria [11][12][13]. Most of these studies indicated that the secondary metabolites produced by Bacillus spp., such as lipopeptides, are low molecular weight substances with constant antagonistic efficacy within broad thermal ranges. In reality, surfactin, which is a bacterial lipopeptide, have detergent like characteristics toward biofilms by reducing the surface tensile strength between these biofilms and adhesion surface [14]. These properties draw researchers’ attention to the possibility of purifying the active phase of bacterial extracts, and their uses as a bio-control agent against harmful biofilm-forming bacteria which are the cause of many environmental problems. Therefore, the aim of this study was to obtain local isolates belonging to Bacillus genus bacteria and to investigate the inhibitory effects of their purified secondary metabolites extracts against the growth of other targeted bacteria native to their environment.
Fifty samples of water and sediment were collected for the period from January 2016 to April 2016 from different places in Basra province (Table 1). The samples were transferred to the laboratory for diagnosis, and then placed in a water bath at 80 °C to kill non-spore forming bacteria [15].
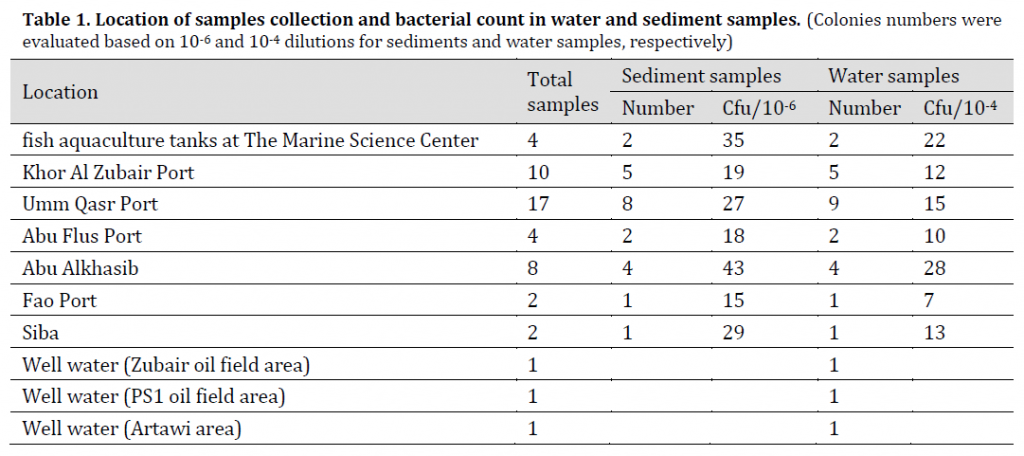
Table 1. Location of samples collection and bacterial count in water and sediment samples. (Colonies numbers were evaluated based on 10-6 and 10-4 dilutions for sediments and water samples, respectively)
Samples were collected (20 g per sample) using sterile dry polyethylene containers. 1 g of each sample was added to a test tube containing 9 ml of distilled water and series of decimal dilutions were created from each sample until 1x10-6 using distilled water [16]. 1x10-1, 1x10-3, 1x10-6 dilutions were filtered using 0.45 µm Millipore filter paper, the filter papers were placed into Luria Bertani (LB) agar petri dishes and the plates were incubated at 35 °C for 18 hr. and diagnosed after incubation.
250 ml water samples were collected using sterile Nalgen polycarbonate conical flasks (Merck, Germany). In sterile conditions, 1 ml of each sample was added to 9 ml of sterile distilled water and this step was repeated to achieve decimal dilution series (1 × 10-1 - 1 × 10-4). After that, 1x10-3 and 1x10-4 dilutions were grown on the same environment for sediment samples and diagnosed [16].
Bacterial colonies of sediment and water samples were inspected under microscope with gram dye and gram-positive spore-forming bacilli were selected for later diagnosis by VITEK® 2 BCL card (bioMérieux, France). Furthermore, bacterial DNA was extracted using Wizard® Genomic DNA Purification Kit, and PCR amplification of the gyrase enzyme gene subunit A (gyrA) was carried out to distinguish Bacillus species [17]. The amplification of (1025 bp) amplicons of the gyrA subunit where used as a differential marker of Bacillus spp. using the following primers:
gyrA-f (5'-CAGTCAGGAAATGCGTACGTCCTT-3') and gyrA-r (5'-CAAGGTAATGCTCCAGGCATTGCT-3') [18].
Polymerase Chain Reaction (PCR) assay was performed using (Applied Biosystem, USA) thermocycler with GoTaq® G2 Green Master Mix (Promega, USA) and according to the manufacturer recommendations. The primers and components were mixed in the same 25 µl amplification reaction containing1µl primer pair mix, 2 µl DNA sample, 12.5 µl Green Master Mix, and 9.5 µl PCR grade water.
The amplification program consisted in a pre-denaturation phase at 94 °C for 2 min followed by 40 cycles: Denaturation 94°C for 30 sec, Annealing 51°C for 45 sec, Extension 72°C for 60 sec and Final Extension 72°C for 7 min. Finally, A 1% agarose gel electrophoresis for PCR reactions were used to distinguish between B. subtilis andother Bacillus spp. bacteria.
Biofilm-forming bacteria were isolated using the specific medium for each bacterial species which are Staphylococcus 110 agar; Pseudomonas isolation agar, Endo agar, Nutrient agar, and Mannitol salt agar then, were biochemically diagnosed using VITEK®GP & GN cards.
Secondary metabolites for Bacillus spp. were produced according to the method of [19]. Bacterial isolates were grown for 48 h at 30°C on a shaker at 120 rpm.
The produced secondary metabolites were extracted. According to [20], Ethyl acetate was added to the bacterial culture in 1: 1 ratio and the mixture was stirred on magnetic stirrer for six hours. After that, the samples were centrifuged at 5000 rpm for 10 minutes. Ethyl acetate layer was collected and dried in a rotary evaporator at 50 °C and the residual was dissolved in ethyl alcohol.
A thin layer chromatography (TLC) assay was performed for the extracted metabolites using silica gel sheets (Silica gel 60 F 254 20 x 20 cm gel thickness: 0.25mm, Merck) with a moving phase consisting of (ethyl alcohol: acetic acid: distilled water) by ratios of (v: v: v 17:38:50) in accordance to [21]. Retention factor (Rf) was calculated using the following formula:
Rf = (distance travelled by the sample) / (distance travelled by solvent)
Total proteins were determined using BIOLABO Total Protein Biuret method kit (Biolabo, France) and following the instructions of the manufacturer. Absorptivity for the samples and standard was recorded using a UV-1800 Spectrophotometer (Shimadzu - Japan) at 550 nm and total proteins in the samples were estimated using the following equation:
Total proteins in the sample = (sample absorbance) / (standard sample absorbance) x standard concentration
Two samples with the highest total protein content were chosen for purification steps using Ion exchange chromatography (IEC) and gel filtration chromatography (GFC) according to [12].
Ion exchange chromatography was carried out according to [12]. (BS8) and (BS14) samples were added to the DEAE cellulose column (25x 3 cm) which was calibrated in phosphate buffer (100 mM, pH7.0). The column was then subjected to three volume wash with the same buffer so unbound proteins separate from the sample. Then, an increased volume of NaCl in the original phosphate buffer (until 100% NaCl) was added to separate the bound proteins from the column. Flow absorbance was continuously observed at 280 nm.
The bound part from the DEAE Cellulose column for each sample was collected and applied to a (Sephadex S-25) column eluted with the same previous phosphate buffer with a flow rate of 1 ml/min and calibrated with potassium dichromate and dextran blue. Absorbance was observed at 280 nm to record absorbance peak [12].
SDS-PAGE for the purified extracts was performed according to [22]. After sample electrophoresis, the gel was stained with Coomassie Blue for a whole night and the dye was removed using a solution of (acetic acid: distilled water: methanol) (4:5:1 v:v:v).
The active molecules in metabolites were determined using GC-MS [23]. An Agilent 78908/5977A GC-MS detector equipped with 30m × 250µm × 0.25µm Hp-5MS capillary column filled with adsorption phase of (5% phenyl, 95% dimethylpolysiloxane). Mass detector was operated on EI mode using helium as carrier gas at a flow rate of 1 ml/min. The injector was operated at 290 °C and the oven temperature was programmed as follows: the initial temperature was 70 °C for 2 minutes and then gradually increased by 16 °C/min until 250 °C. The compounds were diagnosed based on the comparison of their mass spectra with the 2014 NIST Library.
The nature of the chemotaxis between purified secondary metabolites (BS8 and BS14) against targeted bacterial species was tested. The tested targeted bacteria were Pseudomonas aeruginosa, Escherichia coli, Methicillin resistant Staphylococcus aureus (MRSA), Staphylococcus sciuri, and Kocuria kristinae by solid propagation method [24]. In brief, 0.1 ml of liquid bacterial cultures of the previously mentioned species were spread on agar based medium and incubated at 35 °C for 15 min. The medium were pierced with cork-borer and the formed holes were filled with 50 µl of the crude (without purification) or purified metabolite extract (after each purification step). Three replications were carried out for each extract X targeted bacteria combination. Petri dishes were incubated for 18 hours at 35 °C, and inhibition diameters (mm) were then measured using Vernier caliper.
To compare chemotactic effect means, collected data were subjected to analysis of Revised Least Significant Difference (RLSD) using IBM SPSS Statistic software 24 at a probability level of 0.05.
The diagnosis using VITEK® 2 BCL card showed 20 bacterial isolates of the genus Bacillus spp. in the collected samples as shown in (Table 2), the diagnostic results showed that 15 isolates belonged to Bacillus subtilis, while 5 isolates were diagnosed as Bacillus amyloliquefaciens. Furthermore, PCR (gyrA) amplification results were positive in B. subtilis isolates and negative in B. amyloliquefaciens isolates (Fig. 1 A) which refers to the possibility of using this primer pair reported by [17] as an efficient marker to distinguish B. subtilis from B. amyloliquefaciens; however, this method might be limited to some B. amyloliquefaciens strains.
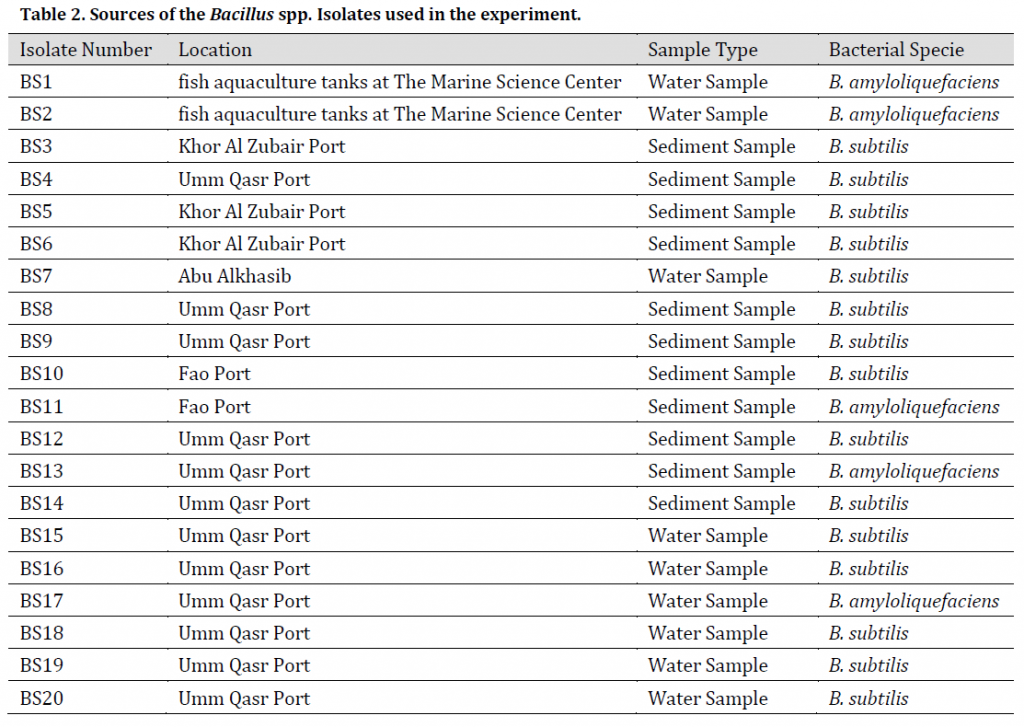
Table 2. Sources of the Bacillus spp. Isolates used in the experiment.
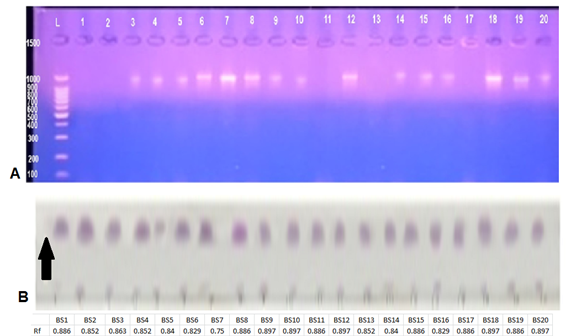

Figure 1. (A) Gel electrophoresis for the gene “gyrA” PCR amplification of bacterial isolates DNA, Positive amplifications of single (1025 bp) bands are noticed for B. subtilis isolates while negative results without amplifications belong to B. amyloliquefaciens isolates, in comparison to L: 1kb DNA ladder (B) Rf results of TLC for secondary metabolites extracts of Bacillus spp. The samples numbered 1-20 represent the 20 different bacterial isolates.
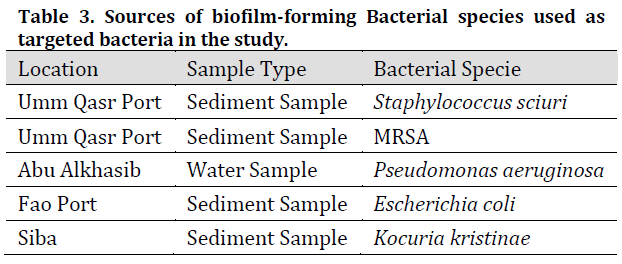
Five biofilm-forming bacterial isolates were diagnosed and isolated using the VITEK GP diagnostic kit for gram-positive isolates and the VITEK GN card for gram-negative isolates respectively and by testing their ability to form biofilms by growth on the specific medium for each bacterial species. (Table 3) shows the results of the diagnosis of target bacterial isolates which agrees with [25] results of these species being able to form biofilms in different environments.
In our study, the selection of secondary metabolite production conditions, especially time, was based on the sheer number of previous studies [26][27][28]. All these works reported that the time required to grow Bacillus spp. species for the purpose of producing secondary metabolites ranged from 24-72 hours. [29] stated that it is 48 hours after growth when the bacterial culture entered the idiophase of stationary phase, and with the decrease of bacterial growth rate, secondary metabolite production begins. This production is usually interpreted as the result of main nutrients consumption such as carbon, nitrogen or phosphorus [30].
In fact, previous studies reported the use of several solvents to extract secondary metabolites from Bacillus bacteria such as N-hexane, dichloromethane, chloroform, methanol, dimethyl sulfoxide (DMSO), and ethyl acetate [23][29][31]. However, secondary metabolites extracted by ethyl acetate showed better antagonistic activity against target Bacillus species compared to metabolites extracted with other organic solvents. This is consistent with the findings in the current study where Bacillus secondary metabolites showed clear inhibitory activity against both negative and positive Gram target species.
The results of thin layer chromatography indicated the appearance of purple spots when using the ninhydrin reagent indicating that the secondary metabolites of Bacillus bacteria contained amino acids as shown in (Fig. 1 B) which also shows Rf values of Bacillus secondary metabolites extracts. These values were similar and ranged between (0.750 - 0.897) which is similar to surfactin Rf (0.7), and in accordance with [32] reports of similar Rf values due to the similarity in molecular weights of extracts.
The results in (Table 4) indicated that the highest value of total proteins in the secondary metabolites extracts belonged to the isolate (BS8) with 13.78 g/l followed by isolate (BS14) with 12.02 g/l while the lowest value was found in (BS19) with 5.28 g/l. Based on these results and TLC results, both (BS8) and (BS14) extracts were chosen for purification and further chemotaxis inspection.
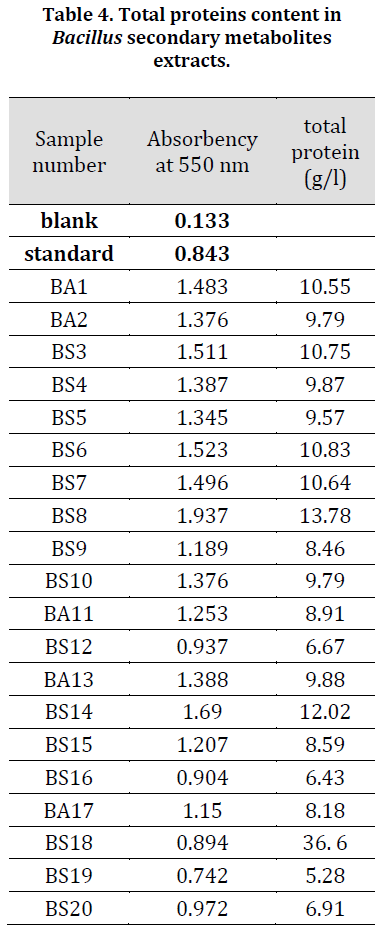
Table 4. Total proteins content in Bacillus secondary metabolites extracts.
It is obvious that IEC for (BS8) and (BS14) samples resulted in two peaked graphs when absorbance collected at 280 nanometres (Fig. 2 A and B respectively). The first IEC peak refers to positive charged and neutral particles, including antimicrobial peptides, which are separated in the first phase, while the second peak refers to the negatively charged particles which are separated in the second phase [12]. IEC stage is crucial in extracts purification due to its high effectiveness in removing the pigments that contaminate the sample and to separate the sample effective phase from the non-effective phase [12][14][33].
GFC was carried out using Sephadex gel S-25, due to the desirable features of this type of gel such as high absorption and separation capacity in addition to its simple preparation procedure and the possibility of re-activation and reuse for several times. GFC for (BS8) and (BS14) samples resulted in a single peaked chart when observed in 280 nm (Fig. 2 C and D respectively) which is consistent with [12][34] results which also referred to the increased inhibitory activity of secondary metabolites GFC purified extracts against target Bacillus species compared to the crude extracts.
SDS-PAGE showed that there was one clear peptide band with molecular weights of 694 Dalton and 3779 for (BS14) and (BS8) purified extract respectively (Fig. 2 E), indicating the efficiency of the methods used in extracting and purifying secondary metabolites. Similar results were obtained by [14] who stated that peptides extracted from Bacillus subtilis appear aggregated in SDS-PAGE gel which is a common characteristic of peptides that possess tensile properties such as surfactin, which has high cleaning and inhibitory abilities against many bacterial species, which qualifies these peptides to be natural alternatives to chemicals used to treat a number of environmental problems, including biofilm-forming bacteria.
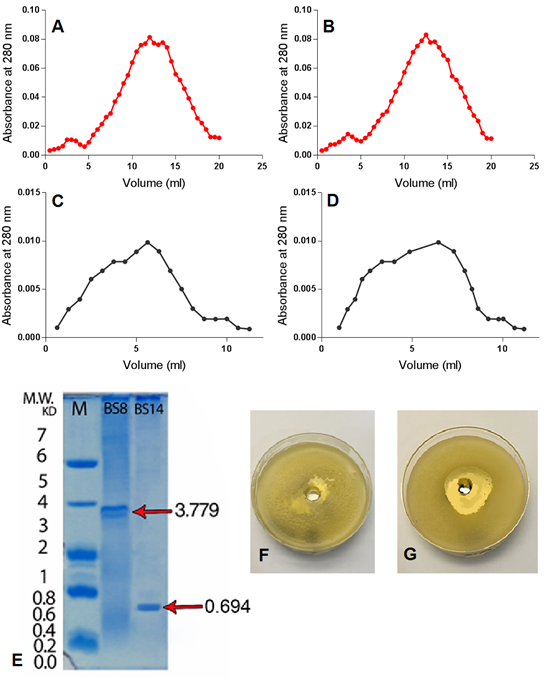
Figure 2. (A) and (B) The elution profile of proteins from the DEAE column eluted with100 mM phosphate buffer (pH 7.0) for (BS8) and (BS14) samples respectively. First peaks represent the unbound (effective) compounds, while the second peaks refer to the bound positively charged (un-effective) compounds. (C) and (D) The elution profiles of purified protein from the Sephadex S-200 column eluted with 100 mM phosphate buffer (pH7.0) for (BS8) and (BS14) samples respectively. (E) SDS-PAGE for the purified extracts compared with an aprotinin polypeptide marker with a molecular weight of 6.512 kDa. (F) and (G) Antimicrobial effect of the purified secondary metabolites extracts (F) BS14 extraction on P. aeruginosa (G) BS8 extract on K. kristinae
GC-MS results showed that purified secondary metabolites extracts include amino acids and their derivatives, esters, hydrocarbons and other compounds (Tables 5 and 6), (TIC scans can be noted in Supplementary material, Supplementary Figs. 1 and 2) many of which have antimicrobial properties such as type D-amino acids [35], N-cyclopropyl carbonyl-[36], and esters (d-proline [37], N-methoxycarbonyl [38], and pentyl ester [39]). Our results are in agreement with study conducted by [40]. In fact, these bioactive extracts were previously described as promising biological control agents for numerous environmental problems caused by biofilm-forming bacteria [41]. Furthermore, type D-amino acids indirectly inhibit the formation of biofilms by interfering with protein synthesis [42]. Interestingly, GC-MS profile for the purified (BS8) extract was more diverse when compared to (BS14) purified extract. We suggest that many metabolites of (BS14) crude extracts were reduced by purification procedure [12] which might explain the few components remaining in GC-MS profile. Additionally, the significant difference between (BS8) and (BS14) GC-MS profiles was previously reported between different bacterial strains of Bacillus and other bacterial species [43][44][45].
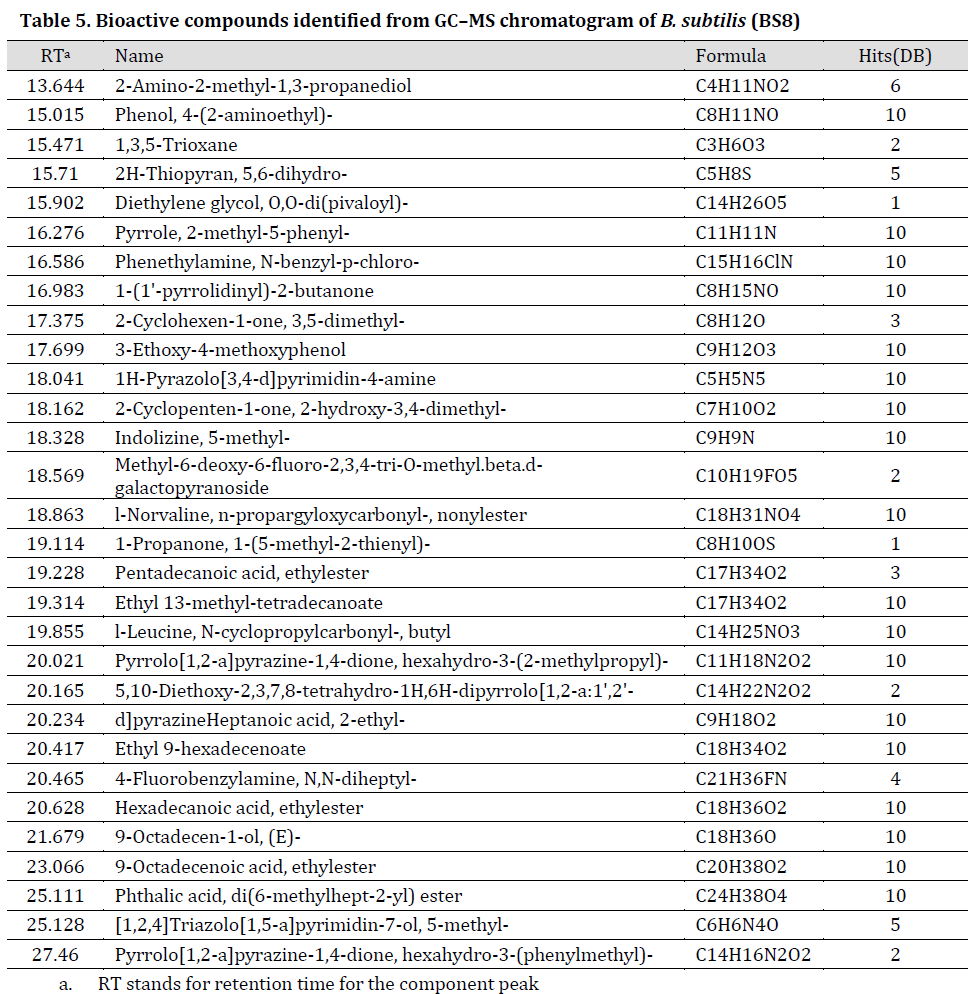
Table 5. Bioactive compounds identified from GC–MS chromatogram of B. subtilis (BS8)
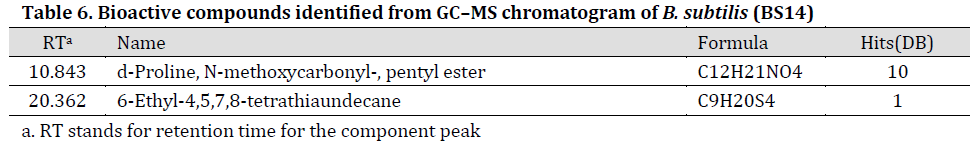

Table 6. Bioactive compounds identified from GC–MS chromatogram of B. subtilis (BS14)
The results showed a clear negative chemotactic effect of the purified secondary metabolites extracts of (BS8) and (BS14) against target bacteria. Transparent halos free of target bacterial growth were formed around the wells containing the secondary metabolites of the isolates. K. kristinae which was the most susceptible species against the purified extracts with 23 and 24 mm for (BS8) and (BS14) extracts respectively, while P. aeruginosa was the least susceptible with 19 and 18 mm for (BS8) and (BS14) extracts respectively (Table 7), (Fig. 2 F and G). Growth inhibition diameters results showed a significant increase in both extracts inhibitory effect after IEC purification and subsequently after GFC purification (Table 7), which indicates the increased efficiency of bacterial extracts using these purification methods. The reaction of targeted bacteria was previously identified as spatial redistribution where the bacteria tends to grow heavily in the places were secondary metabolites were not present [46].
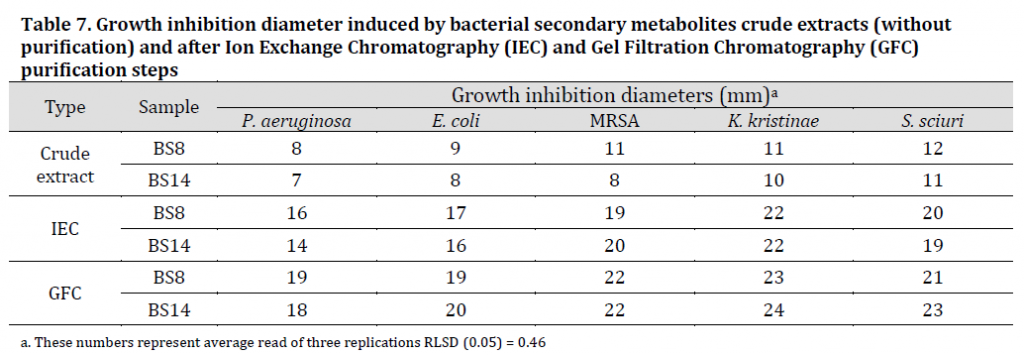
Table 7. Growth inhibition diameter induced by bacterial secondary metabolites crude extracts (without purification) and after Ion Exchange Chromatography (IEC) and Gel Filtration Chromatography (GFC) purification steps
The Iraqi marine environment represents a promising source of bacterial isolates producing various biologically active substances. The rapid growth of bacteria, the ease of growth requirements, and low production costs of bacterial secondary metabolites, might render them as an economic substitute for antibiotics to face the many environmental problems caused by bio-film forming bacteria. Furthermore, a significant increase in bacterial extracts inhibitory by the implementation of simple purification methods such as IEC and GFC.
Conflict of interest statement
The authors declared no conflict of interest.
Funding statement
The authors declared that no funding was received in relation to this manuscript.
Data availability statement
The authors declared that all related data are included in the article and its supplementary material.