

Document Type : Review Article
Author
1 Department of Horticulture, Al-Baath University, Homs, Syria
2 Department of Horticultural Sciences, UTCAN, University of Tehran, Iran
Abstract
Keywords
Artificial intelligence (AI) is defined as the act of thinking and learning when performed by a computer program or machine. Due to its capability of simulating human intelligence and thus performing complicated human tasks, AI is steadily replacing human forces in various sectors. In science, AI is creating new frontiers in environmental [1], medical [2], pharmaceutical [3], biological [4], agricultural [5], and engineering [6] sciences.
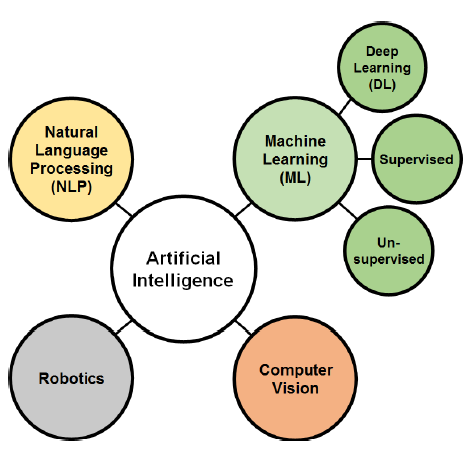
Various methodologies are used in AI research, and classifying these methods is tricky since many of them are interconnected. However, the classification agreed upon by many scholars subdivides AI techniques into four divisions: Machine learning (ML), Natural language processing (NLP), Computer vision, and Robotics (Fig. 1). ML is a major branch of AI that gained increasing importance in the big data era [7]. Deep learning (DL) is a subset field of ML (Fig. 1). The recent rapid growth of DL is generated from the need for a real-time image processing technique, which is the main application of DL through deep and convolutional neural networks (CNN).
Deep learning models are developed to tackle real-life problems in various scientific fields. The aim of this study was to review recent literature of DL applications in plant science. Furthermore, the major difficulties and future aspects of DL in plant science will be discussed.
Deep learning is a comprehensive tool of ML that is constituted upon the principles of artificial neural networks (ANN) with great capability to discover complex features in multilayered data. In fact, the main advantage of DL technique compared to conventional ANNs is its efficiency in automatically extracting the significant features of the analyzed data. This automation allows the researcher to focus on model architecture and results reasoning rather than spending long hours on manual feature extraction [8]. This advantage rendered DL as an ideal tool to tackle many scientific problems in health care [9], bioinformatics [10], finance [11], and agriculture [12].
DL potentials in plant science research are undeniable. The reason behind this certainty is that plant science, similar to all modern sciences, dramatically benefits from visual data and data visualization [13]. Technically, any form of visual input can be used for this technique, whether it is any form of spectral imaging or visualization of data to be interpreted as an image such as metabolic profiles and nucleic acid sequence. However, different DL techniques are suitable for the different data inputs. For instance, CNN models are ideal for high dimensional data such as spectral images inputs, while recurrent neural network (RNN) and long-/short-term memory (LSTM) are preferable for sequential data such as DNA and RNA sequences.
Many approaches can be used to tackle the subject of deep learning applications in plant science. For instance, DL applications can be classified technically based on the DL technique or input data format. However, the current review addresses the subject by classifying DL applications into three distinctive levels: Omics level, Micro/Macroscopic level, and population level.
Omics are a collective field of biological studies that end with the suffix -omics, such as genomics, transcriptomics, proteomics, and metabolomics. These studies address the quantification, structure, and functionality of biological molecules. Therefore, omics provide valuable knowledge of plant organizational functionality [14]. Due to the large data usually provided by omics studies methodologies, DL became a necessary inseparable tool of omics data processing and reasoning.
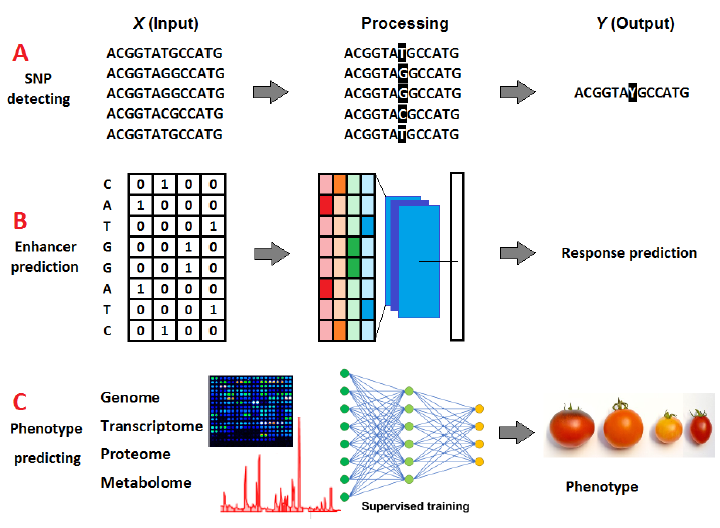
Since omics data is usually sequential, RNNs and LSTM are widely used to process these data. The primary purpose of DL in omics studies is to locate and highlight unique features of interest in the studied data, such as detecting single nuclear polymorphisms (SNPs) (Fig. 2 A) and enhancers’ sequences (Fig. 2 B) [15] or to translate the obtained data (input) into other forms of information (output). For this purpose, molecular data is collectively or individually used to produce a general conceptualization of the plant morphology and phenotypic characteristics, which might have limitless breeding applications [16] (Fig. 2 C).
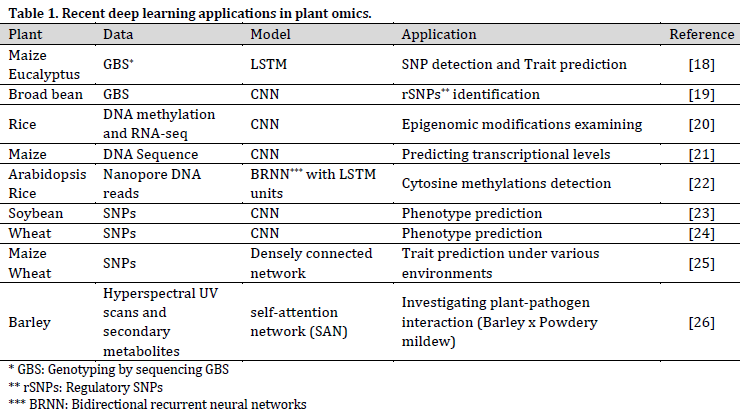
DL techniques are pushing omics research forward in many aspects (Table 1); however, it is stated that other techniques, such as non-additive Gaussian kernel or simple arc-cosine kernel, might produce more reliable predictions based on omics data compared to DL [17]. This observation might be attributed to the overall complicated DL fine-tuning. Therefore, more research should be conducted in the field of DL model’s optimization for omics-based predictions.
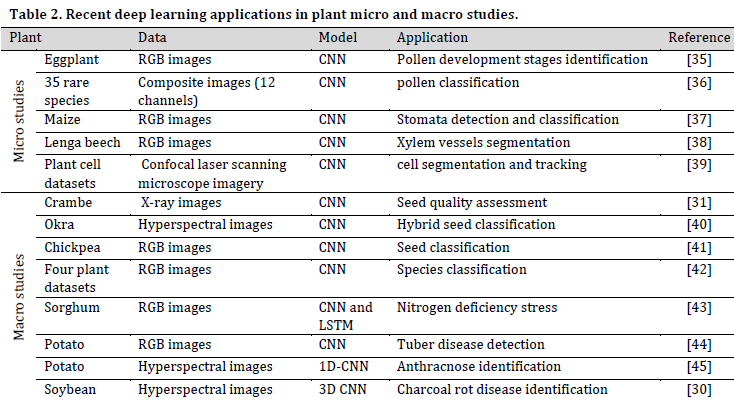
The main concern of microscopic studies is to investigate the plant on a cellular level, such as cellular organelles, full cell, and tissue research. On the other hand, macroscopic studies, in this context, address the characterizations and interactions of a whole or part of a plant. The data used for this type of study are predominantly images of any range of the electromagnetic spectrum.
Various data acquisition systems are being used in Micro/Macroscopic DL, such as visible light sensors [27], infrared (IR) and near-infrared (NIR) [28], ultraviolet [29], hyperspectral [30], and even X-rays [31]. Furthermore, other supportive techniques are being used to increase the depth of the acquired imagery such as cell staining in microscopy [32] and fluorescence imaging systems [33]. There are also some attempts to employ plant’s electrical signals measurements in DL research [34].
The applications of DL in this level are countless such as taxonomy and classification, disease/stress recognition and early warning systems, and physiological events tracking. (Table 2) covers some of the recent studies in the field.
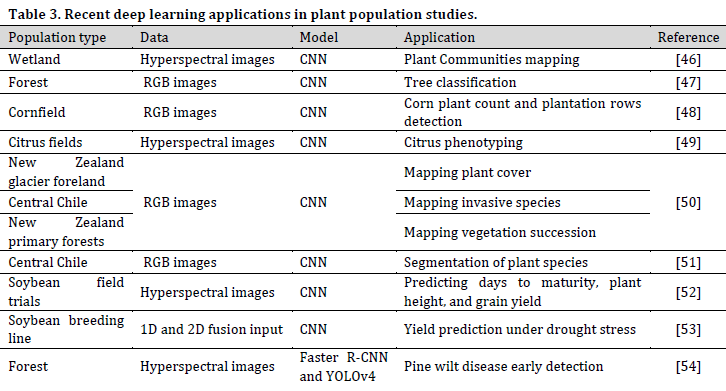
Population plant science refers to the study of plant communities, whether in natural habitats (e.g., forests, grasslands, and deserts) or in agricultural land. Population studies tackle various issues such as plant cover classification and surveys, plant communities’ biological functions, the interactions between individuals or species, and how plant populations are being influenced and influence the surrounding environment (Table 3).
Spatial imagery is the predominant study material in DL population studies. The exponential increase in the availability of remote sensing (RS) datasets increased DL dependant plant population studies. Furthermore, the recent developments in unmanned aerial vehicles (UAV) manufacturing resulted in the production of lighter, more powerful, and affordable drones capable of carrying all sorts of image acquiring systems. These developments created a precious opportunity for research teams to create their RS datasets.
Although DL can provide viable solutions to many complicated issues in plant research, the application of DL in plant science studies is still faced with many obstacles. One of the hurdles facing DL in plant science is dataset size and availability. Due to the relatively high costs of omics research, and the labors related to all three discussed levels of plant research, constructing largely enough datasets is a tiresome task. In fact, DL models depend greatly on the size and balance of the training and validating datasets to illustrate adequate generalization since small datasets usually result in overfitted models [55]. To overcome this obstacle, image augmentation techniques (Fig. 3) are used to expand the dataset size. Various researchers employed data augmentation to increase the accuracy of the developed DL models to some extent. However, increasing dataset size by adding new samples is still required to obtain more reliable results, especially in omics studies, where augmentation techniques are inapplicable.

On the other hand, large datasets require long hours of human supervised labeling since the training process requires adequately labeled data. However, the current rise of self-supervised learning (SSL) and semi-supervised learning might provide a viable solution to this problem since these techniques require unlabeled or partially labeled datasets to learn [56].
Among all three discussed levels of DL research in plant science, omics are still poorly represented. This poor representation is mainly due to the high costs of datasets generating and the unsuitability of omics data in its raw forms to be used in DL training which requires long hours of processing. Therefore, developing new methods for omics data preparation and pre-processing is necessary. Furthermore, employing layer visualizing methods such as saliency map [23] and feature map [27] might be of great importance. These maps provide valuable information regarding the features CNNs use to classify and predict. Therefore, these visualizations can assist omics research in selecting the best plant characteristics.
As for micro/macroscopic levels, it is expected that DL techniques will transform from a complementary assistant tool to play more vital roles. Novel DL models are being developed daily to provide a deeper understanding of physiological processes and biological interactions between plants and their abiotic and biotic surroundings [43-45]. Additionally, DL has excellent potential to provide rapid and accurate judgments in agricultural production lines [27].
Population level DL studies are expected to play significant roles in the real-time tracking of invasive species and plant population dynamics in natural habitats [46][47][50][51]. Furthermore, the new models will introduce a new age of cost-effective and accurate DL-assisted agricultural extension [52][53], which will significantly positively impact food production chains in the near future.
Conflict of interest statement
The author declared no conflict of interest.
Funding statement
The author declared that no funding was received in relation to this manuscript.
Data availability statement
The author declared that all related data are included in the article.