

Document Type : Original Article
Authors
1 Department of Horticulture, Al-Baath University, Homs, Syria
2 Department of Horticultural Sciences, UTCAN, University of Tehran, Iran
3 Department of Animal Science, Ferdowsi University of Mashhad, Mashhad, Iran
Abstract
Keywords
The novel coronavirus (SARS-CoV-2) was originally discovered and spread in Wuhan, China in late 2019 and since then, the disease caused by this β-coronavirus was named by the world health organization (WHO) as coronavirus disease 2019 (COVID-19) [1]. The transmission factors for the disease are still debatable; however, the reservoir of the novel virus is thought to be bats [2] as a huge diversity of coronaviruses is found in them and therefore considered natural reservoirs of SARS-like coronaviruses [3]. While on the other hand, the infectious host from which the virus was originally transmitted to humans is still unknown [1] if there is any.
Many foggy points still surround the outbreak; however, the solid facts are that while the major percentage of COVID-19 cases of had developed mild flu-like symptoms, the rest few suffer from a rapidly developed acute respiratory distress syndrome (ARDS) which might lead to respiratory failure, in addition to multiple organ failure, and even death [4]. Additionally, the long incubation period 3–7 days with no clear evidence of the disease and capability of spreading the infection [5][6] have rendered the virus among the most dangerous especially for older patients and those of major underlining health conditions [1]. Thereafter, and based on the rapid increase in COVID-19 confirmed cases outside of China, WHO announced it a pandemic on 11 March 2020 [7].
SARS-CoV-2 genome encodes different structural and nonstructural proteins. The viral spike (S) protein is the main structural protein in cell invasion process as it facilitates the engagement with angiotensin-converting enzyme 2 (ACE2) of the host cell [8][1] and mediates the viral invasion by priming with host cell Transmembrane protease serine 2 (TMPRSS2) [8] which illustrate the importance of these components in drug design for the pandemic. There are components with the potential of strong binding to the viral spike protein or ACE2 [9][10]. These components could act as molecular blockers for disabling the virus’s ability to enter the cell. On the other hand, the nonstructural main protease (Mpro) facilitates the proteolytic processing of polyproteins and therefore controls the viral gene expression and replication processes which renders it an interesting target for drug design researches [11][12].
Since the original breakout, the efforts to develop an efficient vaccine and pharmaceutical therapies have been rapidly accelerated. Various drugs are already being tested in clinical trials with many showing promising results such as Chloroquine phosphate [13][14][15], Remdesivir [15], in addition to Hydroxychloroquine and Azithromycin [16]. On the other hand, traditional herbal medicine was used before to treat similar breakouts such as SARS and H1N1 which might render herbal extracts and substances as an alternative approach in COVID-19 treatment [17] especially that many of these extracts and compounds are already being used in the treatment of some chronic diseases such as diabetes, HIV/AIDS, and herpes [18][19][20]. The diversity in medicinal plants and their active products demands a rapid evaluation of the possible viral inhibitory effectiveness which might be assisted initially by ligand binding simulations. Therefore, this study aimed to evaluate the binding potentials of various conventional drugs, vitamins, and plant-derived active compounds with SARS-CoV-2 main protease.
The crystal structure of COVID-19 main protease (Mpro) with the ID 6LU7 [12] was retrieved as a PDB file from RCSB protein data bank and imported to Molegro Virtual Docker (MVD) [21]. Taking into consideration the fact that water molecules are not involved in the process of ligand-receptor binding; it is usually preferable to remove them before the molecular docking as this step can significantly enhance the computations and to avoid any probable distortion [22] Water molecules were then deleted and protein molecule was prepared prior to docking simulation via MVD molecule preparation function. Detect cavities function was then used in the search for proper docking constraints on the structure.
In total, 88 conventional antiviral drugs, 16 vitamins, and 63 natural (plant) compounds were chosen from the antivirus lists and literature [18-20][23-26]. All the chosen ligands were downloaded from ZINC15 database in SDF format and then imported into MVD workspace. The ligands were then prepared for docking using MVD molecule preparation function (Fig. 1).
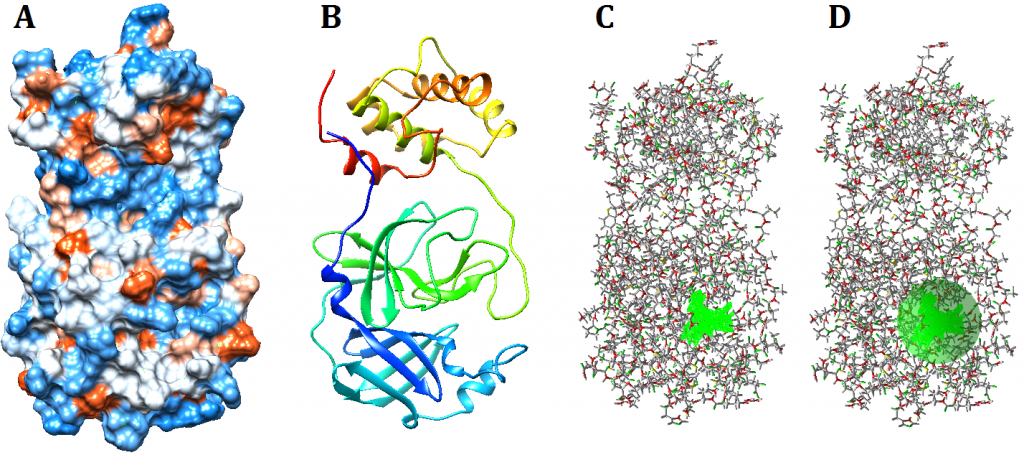
Figure 1. Visualization of SARS-CoV-2 main protease. Visualization using UCSF Chimera hydrophobicity surface (A) and ribbons (B). The chosen cavity (C) which was used to specify docking constraints dimensions and coordinates (D)
MVD docking wizard was started up and the protein structure along with all the chosen ligands were selected for docking. All docking specifications were left as the original default settings; however, binding site constrain was set to include the largest detected cavity on the protein structure and its dimensions were minimized in order to decrease the simulation processing time and increase accuracy (Fig. 1 D). Furthermore, the number of runs was set to 50 runs/ligand and the maximum population was set to 100 according to MVD recommendations as these settings resulted in better outcomes.
After completion, docking results were imported and the best pose of each ligand, based on their Re-rank score, was loaded into the workspace. The 2D diagrams of receptor-ligand interaction for the best poses were visualized using BIOVIA discovery space visualizer; while docking poses within the protein structure were visualized using UCSF Chimera software after exporting the protein molecule and poses in MOL2 format.
Cavity prediction function results demonstrated the presence of four cavities on the surface of Mpro which ranged in volume from 10.24 – 126.98 Å. The largest cavity was chosen for docking as it has originally included the biding ligand 02J-ALA-VAL-LEU-PJE-010 as provided in PDB (Fig. 1 C).
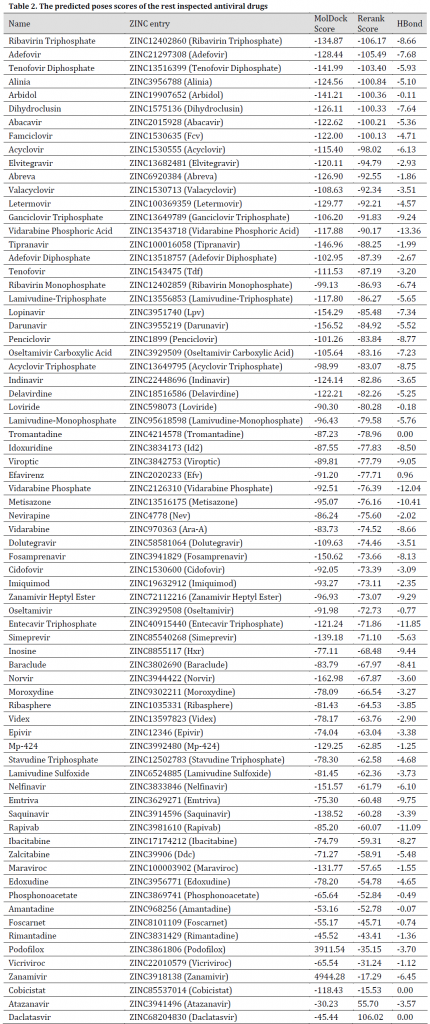
All the chosen compounds were evaluated for binding probability in the same docking session. After docking, the molecules were ranked based on their Re-rank score. The results showed that the best drugs were Tenofovir Alafenamide, Adefovir dipivoxil, Amprenavir, Raltegravir, Intelence, Ganciclovir, Ganciclovir Triphosphate, Pleconaril, Acyclovir Diphosphate, Doravirine, Rilpivirine, Valganciclovir, Victrelis, Stavudine, and Tenofovir Disoproxil with a re-rank scores ranged from -117.493 to -106.829 and MolDock scores between -167.369 and -125.673 (Table 1). Additionally, graphical visualization showed strong hydrogen bonds interaction with protein molecule (Fig. 2-6). All previously mentioned drugs are used in treatment programs of various viral diseases including HIV/AIDS, Cytomegalovirus (CMV), herpes, and Picornavirus (Table 1). Although some of these drugs have entered clinical trials of COVID-19, there are some reports of few with no significant positive results such as Ganciclovir and Acyclovir (no reports of Acyclovir Diphosphate) and therefore, these drugs were not recommended [1][44][45], additionally, Tenofovir Alafenamide trials [46] have also started with no new reports so far. The rest simulated drugs result can be seen in (Table 2).
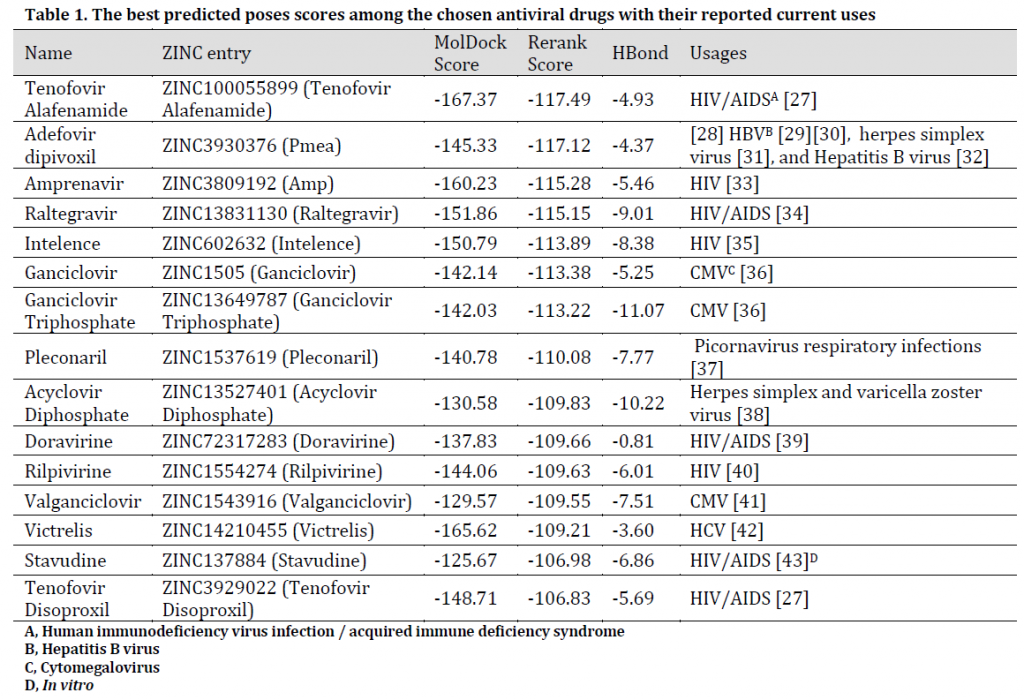
Table 1. The best predicted poses scores among the chosen antiviral drugs with their reported current uses
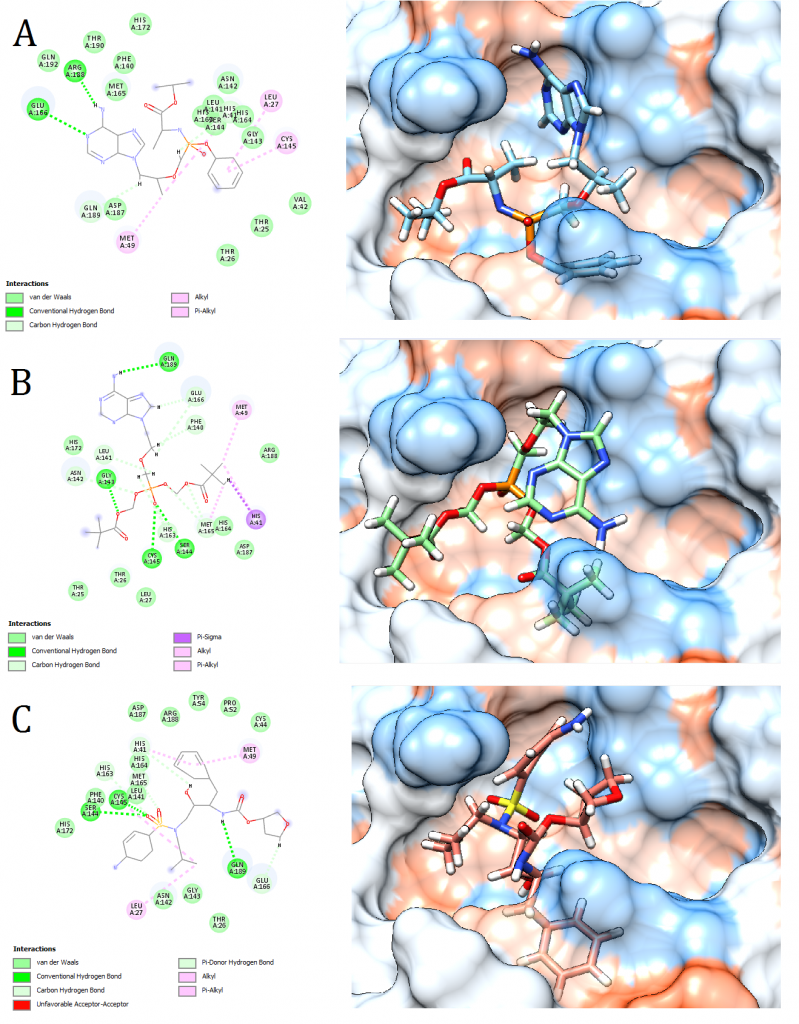
Figure 2. Docking pose visualization for the drugs with the best predicted results (Right) and their 2D receptor-ligand interaction diagram (Left). (A) Tenofovir Alafenamide, (B) Adefovir dipivoxil, (C) Amprenavir
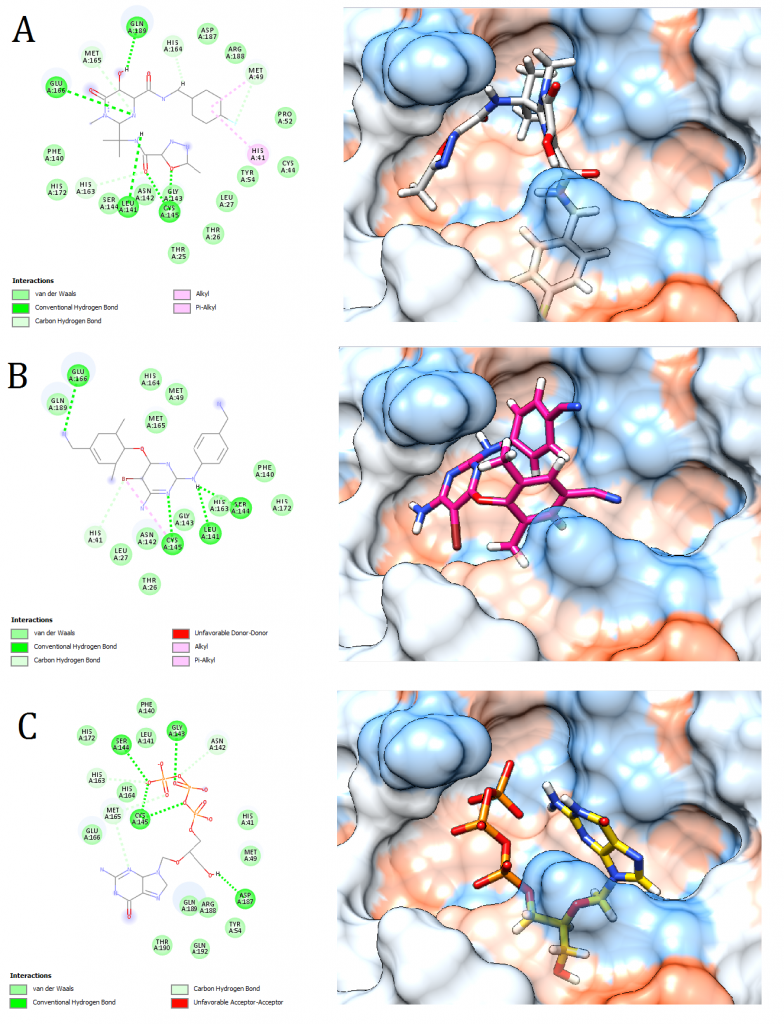
Figure 3. Docking pose visualization for the drugs with the best predicted results (Right) and their 2D receptor-ligand interaction diagram (Left). (A) Raltegravir, (B) Intelence, (C) Ganciclovir
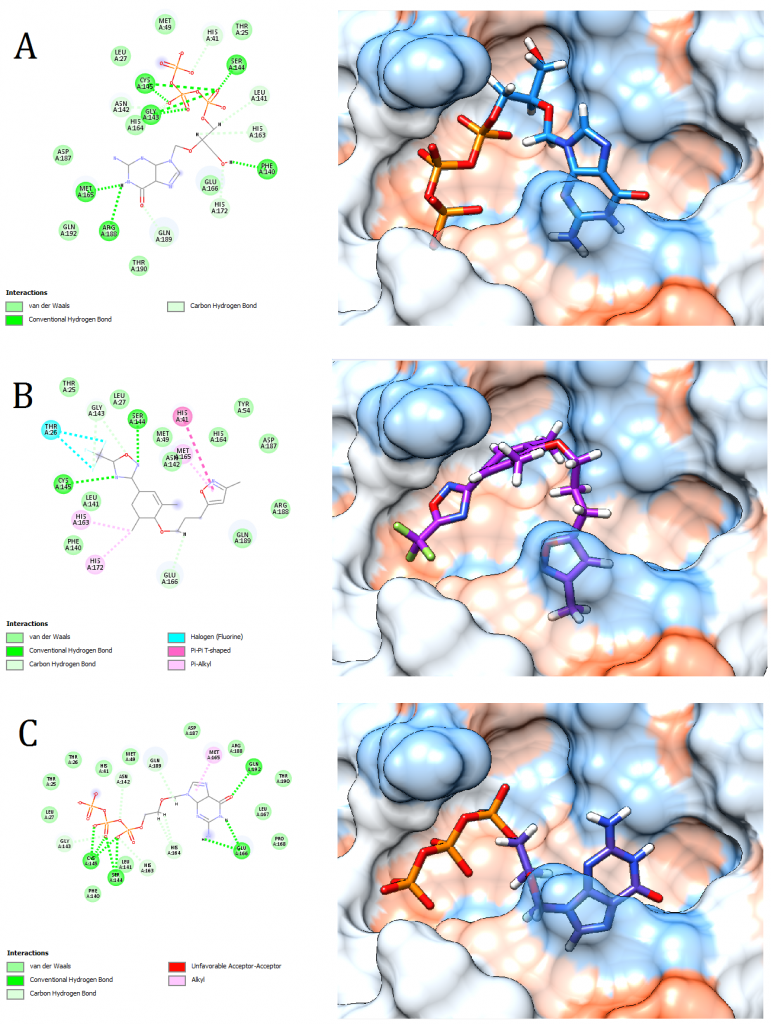
Figure 4. Docking pose visualization for the drugs with the best predicted results (Right) and their 2D receptor-ligand interaction diagram (Left). (A) Ganciclovir Triphosphate, (B) Pleconaril, (C) Acyclovir Diphosphate
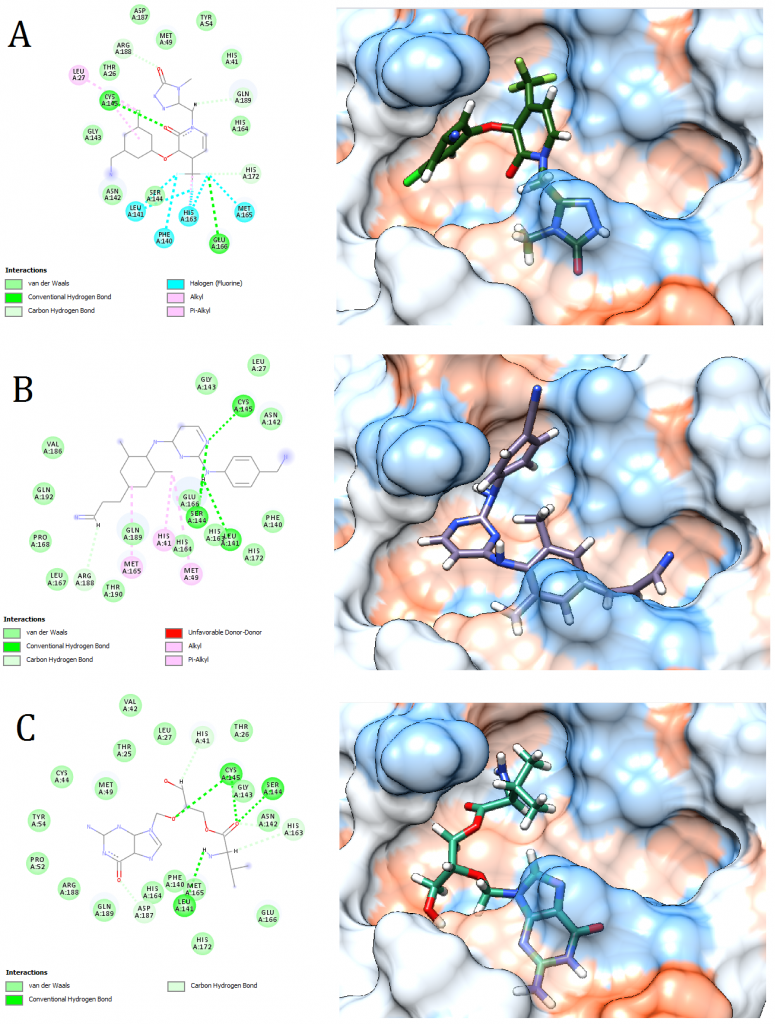
Figure 5. Docking pose visualization for the drugs with the best predicted results (Right) and their 2D receptor-ligand interaction diagram (Left). (A) Doravirine, (B) Rilpivirine, (C) Valganciclovir
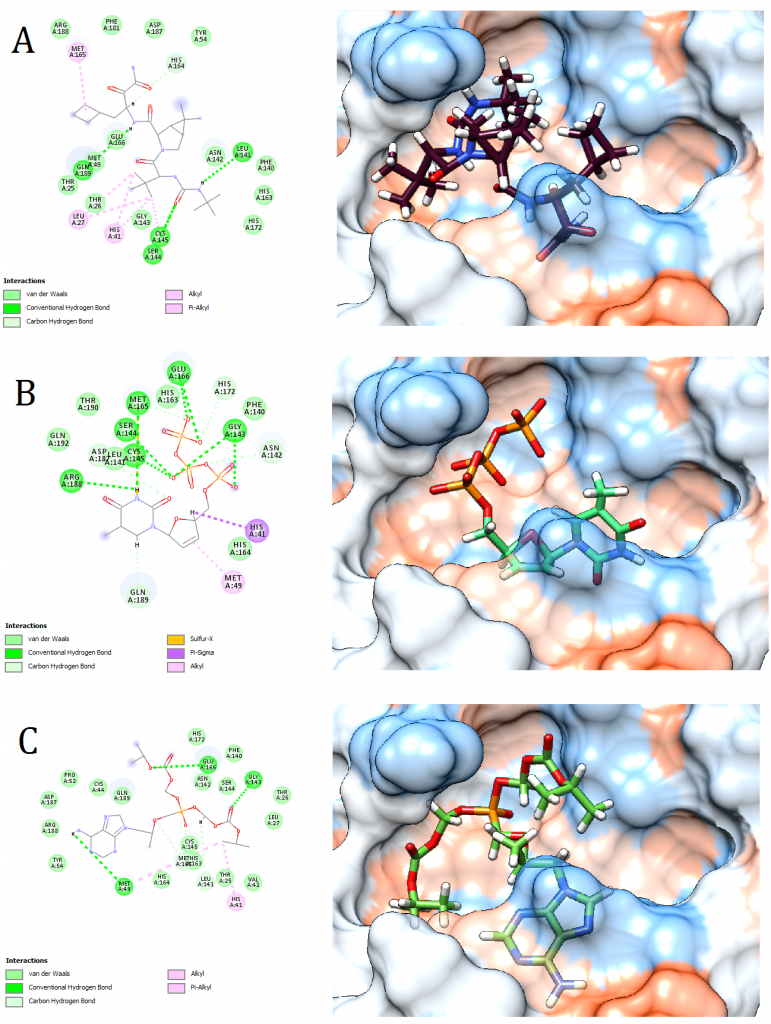
Figure 6. Docking pose visualization for the drugs with the best predicted results (Right) and their 2D receptor-ligand interaction diagram (Left). (A) Victrelis, (B) Stavudine, (C) Tenofovir Disoproxil
Among the investigated vitamins, B9, A, K, and E vitamins exhibited promising results with MolDock scores between -131.99 and -148.43 (Table 3). It is always recommended to fortify body immunity through a balanced diet that satisfies the daily body needs of vitamins and minerals. Vitamin B9 was additionally reported to have an Inhibitory effect on RNA-dependent-RNA polymerase activity of SARS-CoV-2 virus’s SCV2-nsp12 enzyme. On the other hand, Vitamin K3 was reported to have an inhibitory effect on HIV [47]. Additionally, vitamins E and D were reported to be proper candidates in improving body immunity towards COVID-19 [48], as low levels of these supplements increased viral infections with bovine coronavirus in cattle [49].
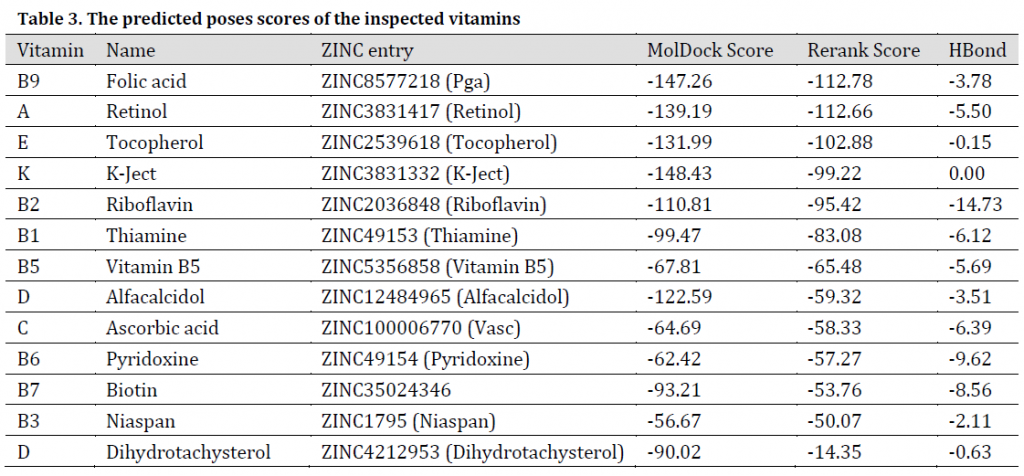
Table 3. The predicted poses scores of the inspected vitamins
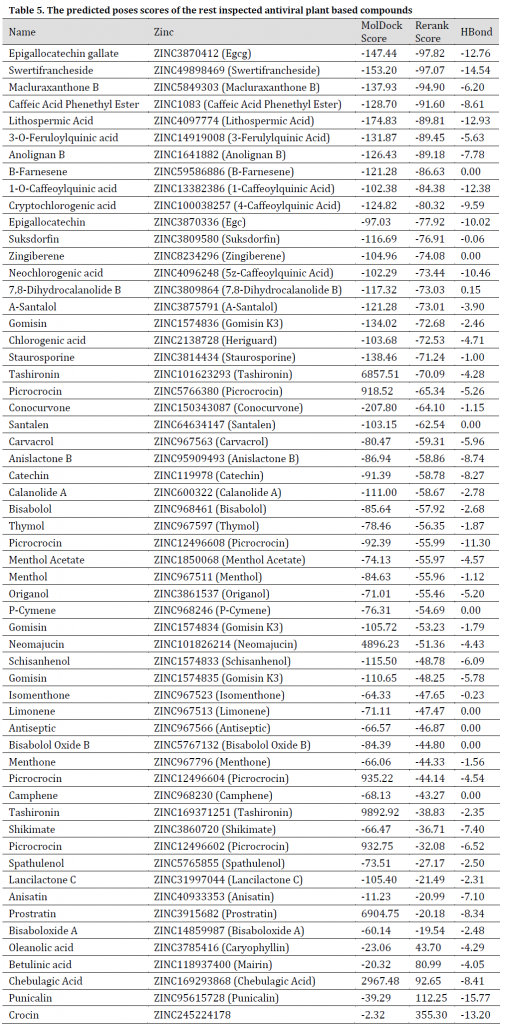
As for the studied plant natural compounds, five substances showed promising results. These substances were Curcumin (Curcuma longa), Mallotojaponin (Phyllanthus myrtifolius), Peltatol A (Pothomorphe peltata), Anolignan A (Anogeissus acuminata), and Phyllamyricin B (Phyllanthus myrtifolius) with rerank scores ranged between -131.59 and -107.42 with MolDock scores between -219.39 and -156.27 (Table 4) and various formed conventional Hydrogen bonds (Fig. 7 and 8). All these substances were reported as HIV inhibitors [24][50-54]. Curcumin was previously reported as a strong SARS main protease inhibitor [55]. The rest of the investigated natural-based molecules can be seen in (Table 5).
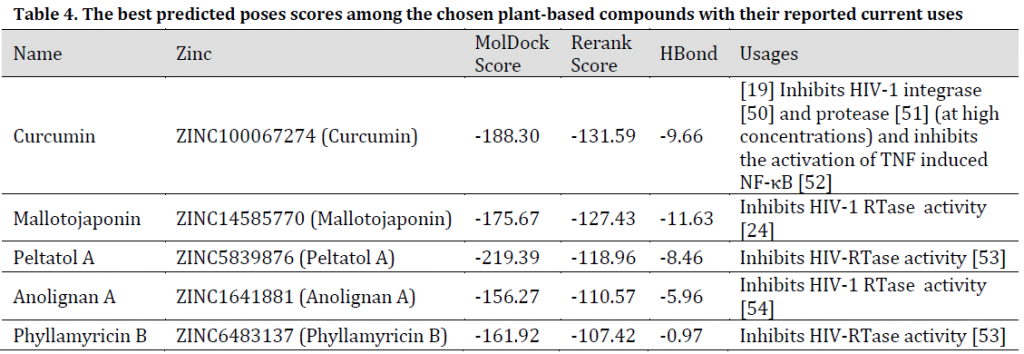
Table 4. The best predicted poses scores among the chosen plant-based compounds with their reported current uses
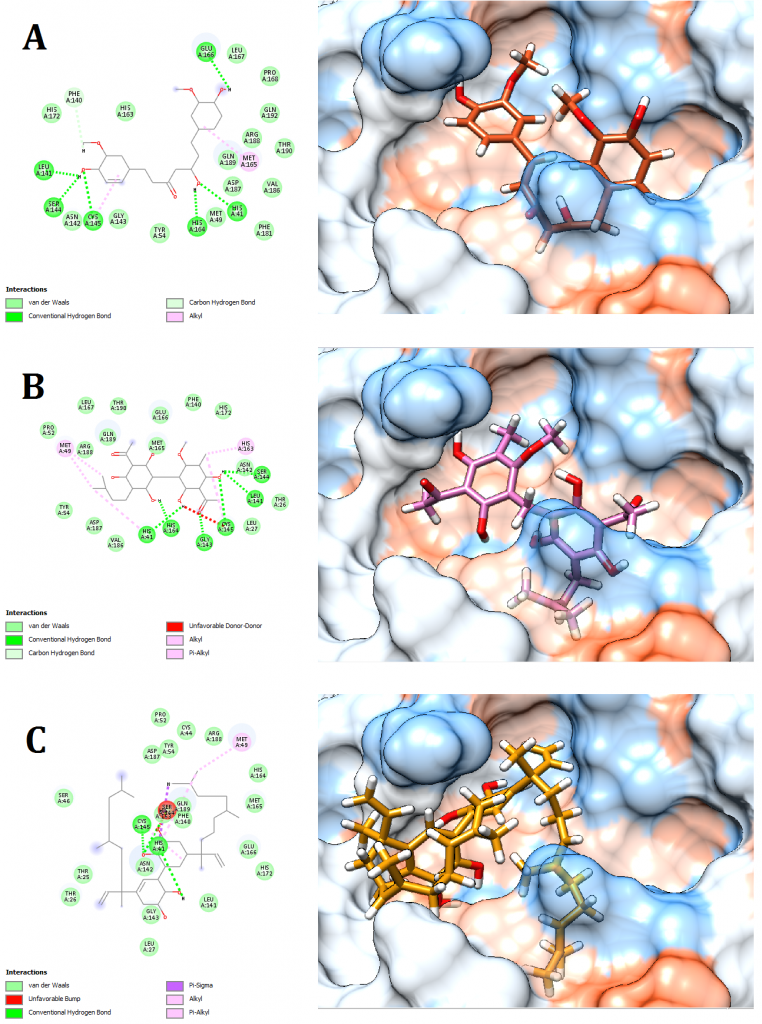
Figure 7. Docking pose visualization for the plant compounds with the best predicted results (Right) and their 2D receptor-ligand interaction diagram (Left). (A) Curcumin (Curcuma longa), (B) Mallotojaponin (Phyllanthus myrtifolius), (C) Peltatol A (Pothomorphe peltata)
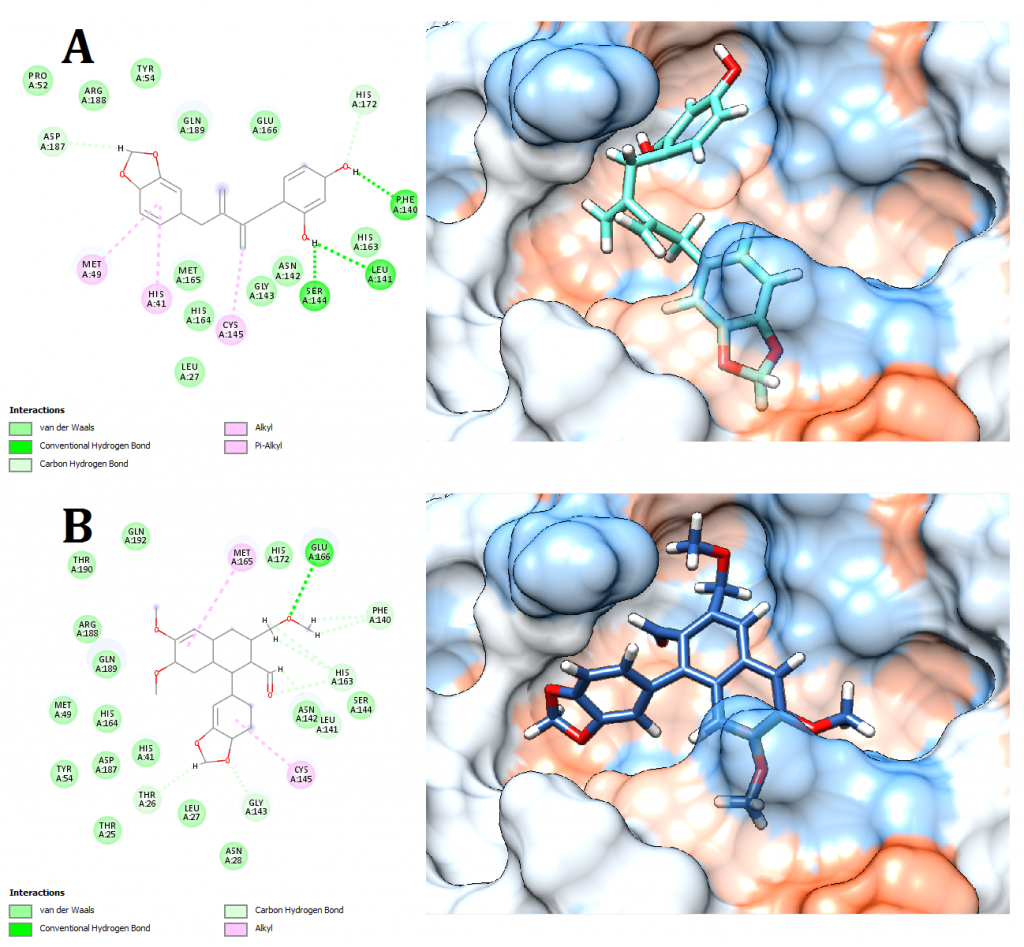
Figure 8. Docking pose visualization for the plant compounds with the best predicted results (Right) and their 2D receptor-ligand interaction diagram (Left). (A) Anolignan A (Anogeissus acuminata), (B) Phyllamyricin B (Phyllanthus myrtifolius)
2D receptor-ligand interaction diagrams illustrated that among the 15 best drugs; Amprenavir, Intelence, Ganciclovir, Acyclovir Diphosphate, and Rilpivirine had an unfavorable acceptor-acceptor bond as can be seen in (Fig. 2 C), (Fig. 3 B), (Fig. 3 C), (Fig. 4 C), and (Fig. 5 B). On the other hand, among the 5 best natural products, Mallotojaponin had an unfavorable acceptor-acceptor bond (Fig. 7 B) while Peltatol A had an unfavorable bump (Fig. 7 C). Ganciclovir and Acyclovir were previously reported to have no inhibitory effect on the novel virus and were therefore not recommended for treatment which might be partially correlated to the predicted unfavorable bonds illustrated in the current study. However, in addition to their various conventional hydrogen bonds; the rest shortlisted drugs (except for Ganciclovir Triphosphate and Valganciclovir) and plant compounds were found to have various Alkyl and/or Pi-Alkyl bonds which might positively contribute to the hydrophobic ligand-receptor interaction [56]. Pi-Pi and fluorine bonding which can be seen in the interaction diagram of Pleconaril (Fig. 4 B) play an essential role in protein-ligand binding [57] and might render the binding of this drug as one of the strongest among the tested ligands [56]. Charge transfer which might be involved with Pi-sigma bonds present in Adefovir dipivoxil-protein and Stavudine-protein interaction diagrams (Fig. 2 B) and (Fig. 6 B) respectively can support an intercalating binding between these drugs and the receptor [56].
COVID-19 is without a doubt a crisis that grew to a global scale affecting the lives of billions worldwide. Therefore, all the efforts that may lead to a better understanding or provide significant relief of the major symptoms are highly welcomed. The results of the current study emphasize on the importance of vitamins in daily diets for their probable inhibitory effect on the virus especially B9, A, K, and E vitamins. Additionally, various conventional drugs and plant-based compounds have shown interesting ligand-protein interactions and therefore worth further investigation especially Pleconaril, Adefovir dipivoxil, and Stavudine in addition to Curcumin (Curcuma longa), Anolignan A (Anogeissus acuminata), and Phyllamyricin B (Phyllanthus myrtifolius). Molecular blockers for the virus or cell surface receptors were not investigated in the current study; however, these approaches might find other alternative potential drugs for this global epidemic.
Conflict of interest statement
The authors declared no conflict of interest.
Funding statement
The authors declared that no funding was received in relation to this manuscript.
Data availability statement
The authors declared that all related data are included in the article.